1. What is Proteomics?
Proteins are essential bio-macromolecules that play a vital role in all living organisms. They perform a variety of functions, from forming structures such as muscle fibers, catalyzing biochemical reactions (through enzymes) to participate in DNA synthesis and replication.
The entire set of proteins potentially expressed by a genome, cell, tissue, or organism at a given time is called the proteome. The proteome is the functional foundation of a cell and organism. The term 'proteome' is a combination of 'protein' and 'genome'.
Proteomics is the scientific discipline dedicates to the study of the proteome. Proteomics researchers focus on determining the identity, structure, abundance, and interactions of proteins. The primary objective of proteomics is to gain the deepr understanding of the role of proteins play in biological processes, enabling applications in disease research, drug development, and various other domains.
2. Proteomic Analysis
Proteomic research primarily employs two approaches: top-down and bottom-up.
The bottom-up approach, also known as peptide-based proteomics, involves enzymatically digesting proteins into peptides, typically using trypsin. These peptides are then separated through specialized chromatographic columns and analyzed via mass spectrometry (MS). The bottom-up approach can be categorized into two groups based on the separation step:
- Two-Dimensional Gel Electrophoresis (2-DE): Proteins are first separated from a gel, followed by enzymatic digestion into peptides, which are subsequently identified using MS
- Shotgun Proteomics: Protein are digested into peptides prior to separation, and liquid chromatography (LC) is used to separate peptides before MS analysis.
In top-down proteomics, intact proteins or large polypeptides are directly analyzed by MS. The molecular weight of these proteins is typically determined using electrospray ionization (ESI) followed by matrix-assisted laser desorption/ionization (MALDI) MS. This method enables the identification of proteins with molecular weights exceeding 200 kDa.
Protein Characterization
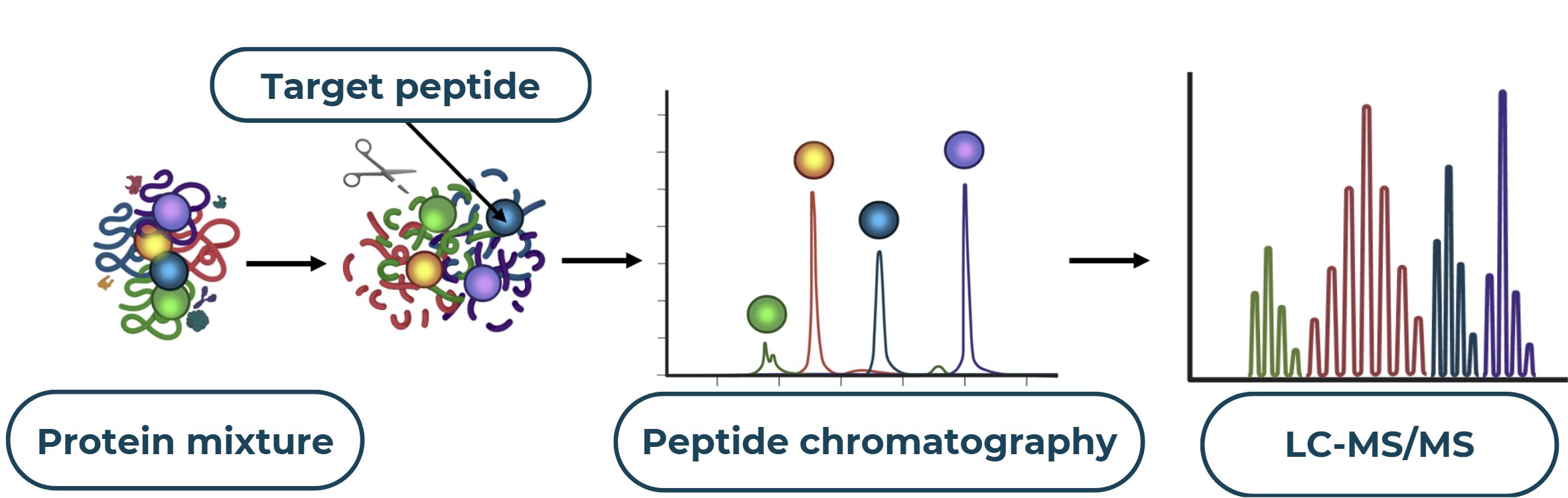
The process of protein analysis begins with the complex proteins are broken down into peptides using specific enzymes. Subsequently, these peptides are separated based on their size and charge through chromatographic or electrophoretic techniques. Each peptide is then ionized and analyzed by MS to determine its precise mass and amino acid sequence. Finally, the obtained information is compared against known protein databases to identify the original protein.
Targeted Proteomics

A typical quantitative protein analysis workflow follows these five steps: sample preparation è protein digestion è peptide separation using high-performance liquid chromatography (HPLC) è peptide ionization and fragmentation utilizing mass spectrometry (MS) è data processing.
After proteins are hydrolyzed into peptides by specific enzymes, the resulting peptide mixture is separated on an HPLC column using an appropriate mobile phase. The target peptides are introduced into MS for ionization using electrospray ionization (ESI) or matrix-assisted laser desorption/ionization (MALDI) technique. The peptide ions are then fragmented to generate ions, from which the amino acid sequence and molecular mass of the peptide can be determined. The acquired MS data is processed using specialized software to accurately quantify peptides using methods such as multiple reaction monitoring (MRM) or parallel reaction monitoring (PRM).

